Mechanistic Evolutionary Biology using the Awesome Power of Non-model Yeasts
Centromere Structure | Cell Cycle Division | Chromatin | Evolution | Genetics | Biochemistry
Current Research
My current research explores how centromeres evolve and how centromeres nucleate kinetochore assembly. I use multidisciplinary approaches such as phylogenetics, genomics, biochemistry, and structural methods. I use yeasts as a model system and look to make Saccharomycotina a model subphylum for cell biology.

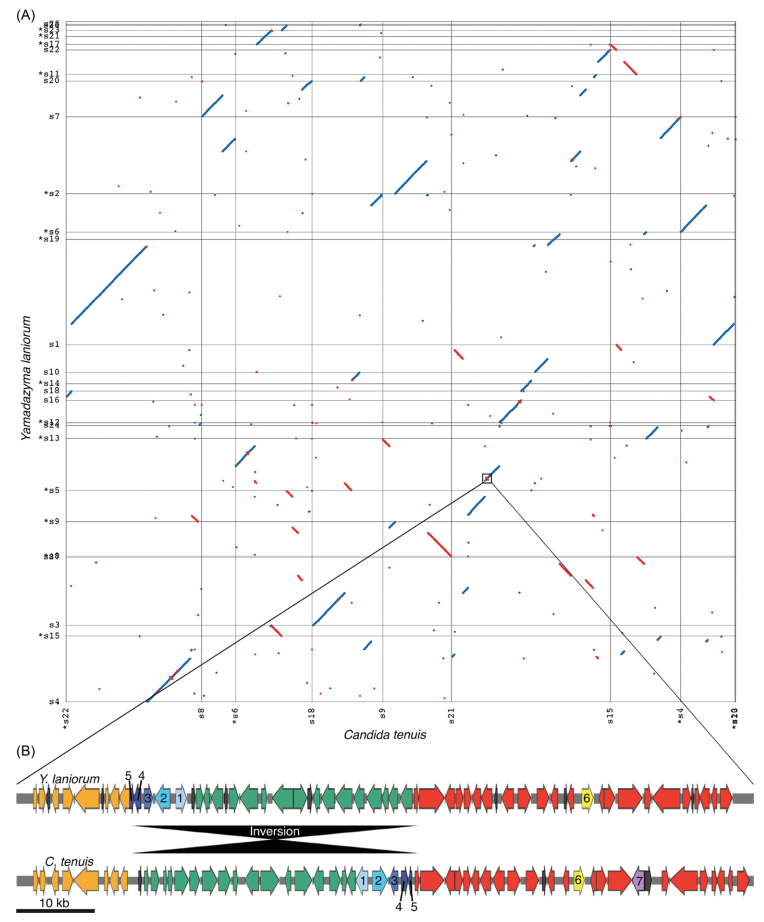
Unlike most eukaryotes, brewer’s yeast have genetic “point” centromeres. Despite being the first centromeres discovered over forty years ago, we still have little understanding of how they evolved. Because point centromeres share no clear homology with other centromere types, it has been difficult to reconstruct their origins.
In our work on a sister group to brewer’s yeast, we identified a new class of centromere—“proto-points”—that appears to represent an evolutionary transitional state. These findings suggest that point centromeres are direct descendants of LTR retrotransposons.
I’m also interested in how historical contingency shapes modern centromere function. From this perspective, I aim to understand how the ancient allopolyploid whole-genome duplication event influenced the present-day function of S. cerevisiae centromeres. I first presented this observation and hypothesis during my PhD work.
Representative publications (@ = Corresponding Author)
Despite our recent progress in understanding how yeast centromeres evolved, we still lack a detailed molecular view of how centromeres nucleate kinetochore assembly. I am now studying native centromere–kinetochore structures to reveal, at atomic resolution, how centromere DNA drives this process.
Other Efforts
Deep Mutational Scanning
During my PhD, I worked independently with the Bhabha and Ekiert labs at Johns Hopkins University to develop a pipeline for deep mutational scanning of bacterial phospholipid transporters (e.g., MlaC, LetA). I have continued this work with Priyanka Verma at Washington University.
Representative publications
Gallery
Below are some videos and images taken from my research.